Objectives of Proposed Research
- To survey and molecularly identify Fusarium oxysporum f. sp. vasinfectum (FOV) races and other seedling and wilt pathogens in commercial and grower cotton fields in California.
- To further evaluate the seedling and wilt capabilities of FOV races with different inoculation methods using susceptible and resistant Pima and Upland germplasm.
- To further evaluate the interactions of different FOV races and Rhizoctonia solani and their impact on disease development in cotton.
Additional match funding has been approved from the California State University Agricultural Research Institute (ARI) for both years of the project thanks to support letters provided by CCGGA and Cotton Inc. With the additional funding, we were able to expand on our current proposed work and add the following objectives to the ARI proposal.
- To use representative identified FOV races for phenotypic evaluation of selected Upland cotton germplasm
- To determine the effects of pH, temperature, and moisture on disease development in cotton when inoculated with FOV4.
Objective 1: To survey and molecularly identify Fusarium oxysporum f. sp. vasinfectum (FOV) races and other seedling and wilt pathogens in commercial and grower cotton fields in California.
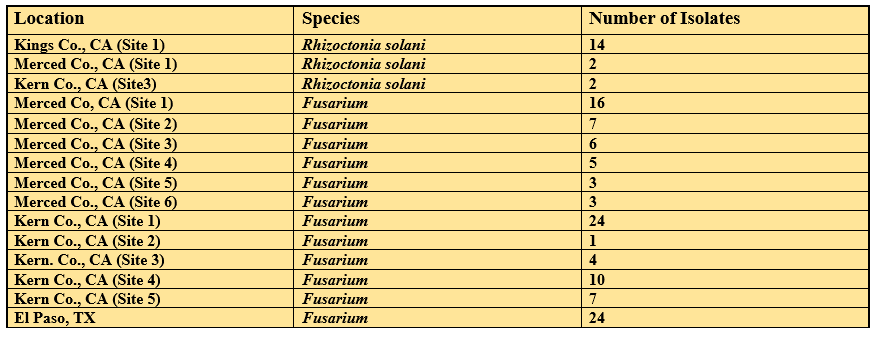
Prior to this proposal, Fusarium isolates were collected in 2017 and 2018. Isolate information is provided in Table 1. All isolates were identified using two PCR assays and DNA sequencing of the translation elongation factor (EF-1α) gene. The first PCR assay produced a 208 bp amplicon unique to FOV races 3, 4, and 7, while the second multiplex PCR assay genotyped FOV isolates into two genotypes, N (396 bp), and T (583 bp). These genotypes were identified based on the absence (N type) or presence (T type) of the insertion of the transposable element Tfo1 in the phosphate permease (PHO) gene unique to FOV race 4. Although not shown these isolates have been genotyped with newly developed primers. We are repeating the genotyping currently for verification of results.

For the current proposed research, 11 locations across the San Joaquin Valley and six locations in the El Paso, Texas region were sampled beginning in mid-May 2019. To date, single spore isolations for 110 Fusarium isolates have been completed (Table 2). All isolates were collected from symptomatic cotton seedlings. Additionally, 18 isolates of Rhizoctonia solani were also isolated from symptomatic cotton seedlings (Table 2). Other fungal species were isolated and are currently being identified morphologically. It appears that there may be some additional Fusarium species that are not FOV. Isolates will be genotyped similar to 2017 and 2018 isolates.
A preliminary baiting method using collected soil from a cotton field in Dos Palos, Calif. was completed. This assay was modified for the isolation of Pythium spp. from field soil using soybean as bait. From this assay isolates of what appear to be FOV, Pythium, and R. solani were all baited using the susceptible Pima cultivar DP-340. Soil has been collected from a number of locations across CA and will be used in the baiting method to isolate other potential pathogens not isolated from collected plant material. Soil samples from some of the first locations where FOV race 4 was identified in CA but are no longer in production for cotton were also collected. This assay might allow us to determine if the pathogen FOV race 4 is still present in these locations, despite being out of production for at least a decade in some cases.
Two undergraduate students have been trained and have been conducting the work mentioned above under the guidance of Dr. Ellis and her previous graduate student. Another student has also started to isolate DNA from the single spore isolations for identification using new PCR primers. Additionally, DNA sequencing of isolates will be done using the translation elongation factor and internal transcribed spacer region.
Objective 2: To further evaluate the seedling and wilt capabilities of FOV races with different inoculation methods using susceptible and resistant Pima and Upland germplasm.
Three assays will be compared to further evaluate seedling and wilt capabilities of FOV races/genotypes. A rolled towel assay was developed in our lab, and will be compared to the root-dip inoculation method and an infested-oat-seed method that was modified from a protocol by Beccera et al. (2012). Protocols for these methods have all been established and tested in preliminary studies. A rolled towel assay using eight representative Fusarium isolates was completed to evaluate possible variation in aggressiveness towards cotton by different FOV4 genotypes and F. solani isolates. The assay was set up using Pima cultivar DP-340. The results from two runs of the assay are provided in Figure 1 and 2, below. There was a significant difference among isolate and experiment (P<0.0001), but there was not a significant difference for the interaction for isolate and experiment.
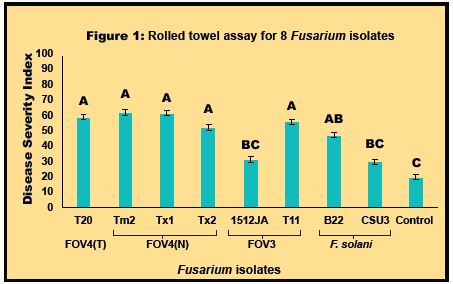

Additionally, these same isolates or a similar set will be used in the comparison of different greenhouse assays. We plan to use varieties of both Pima and Upland cotton with varying levels of plant host resistance in the assays. Finally, we have started to screen previous isolates collected from 2017 and 2018 using the root dip inoculation method. This will also be done for a majority of isolates collected in 2019. Once pathogenicity for the majority of the isolates is tested using the root dip assay and genotyping is completed a representative set of isolates can be used in our screening efforts.
Objective 3: To further evaluate the interactions of different FOV races and Rhizoctonia solani and their impact on disease development in cotton.
The graduate student for this objective has been currently evaluating environmental parameters of our CA FOV and R. solani isolates such as pH and temperature. Infested-oat inoculum has been prepared to begin the interaction study with FOV race 4 and R. solani. Furthermore, we also plan to co-inoculate with different FOV race 4 genotypes and F. solani.




















